|
The pattern formed by the Min proteins allows an E. One widely studied pattern is made in a rod-shaped bacterium called Escherichia coli by a group of proteins called the Min proteins. Some proteins can spontaneously organize themselves into ordered patterns within living cells. Our results provide an essential experimental base for modeling of intracellular Min gradients in bacterial cell division as well as, more generally, for understanding pattern formation in reaction-diffusion systems. Confinement as well as an elevated temperature reduce the characteristic wavelength of the Min patterns, although even for confined chambers with a bacterial-level viscosity, the patterns retain a ~5 times larger wavelength than in vivo. We establish the geometrical selection rules and show that, surprisingly, Min-protein spiral rotations govern the larger part of the geometrical phase diagram. We identify three typical dynamical behaviors that occur dependent on the geometrical chamber parameters: pole-to-pole oscillations, spiral rotations, and traveling waves. Here we present the first in vitro study of the Min system in fully confined three-dimensional chambers that are lithography-defined, lipid-bilayer coated and isolated through pressure valves. The bacterial Min protein system provides a major model system for studying reaction-diffusion processes in biology. Scale bar for the kymographs are shown next to panel ( h). ( e– h) Corresponding kymographs of the MinD intensity along the cells’ length for panels ( a– d) respectively. For the last case, instead of the regular pattern, where triple-point-oscillations occur between a middle zone and the two poles (see panels ( b and f)), the middle zone seems to originate in the center of the cell and while traveling toward one pole, a MinD zone is established on the opposite pole. ( d) Aberrant triple-node oscillations in a Cephalexin-division-inhibited cell (14.4 μm). ( c) Multiple-node oscillations in a Cephalexin-division-inhibited cell (19.9 μm). ( b) A triple-node oscillations in a Cephalexin-division-inhibited cell with a length of 11.9 μm. ( a) Pole-to-pole oscillations in a wild type cell of normal length (3.8 μm). Scale bar (5 μm) is shown to the left of panel (d). Panels show montages of the corresponding supporting Supplementary file 2. ( a– d) Four examples of in vivo MinD oscillations in cells with various length of strain FW1919 (see Wu et al., 2016) Mol Sys Biol 12: 873). This dialog is activated by the Set Column Values.Supplementary file 1: Examples of in vivo oscillations in live cells. It allows to fill a column with the result of a function. 
The available mathematical functions (assuming you are using the default scripting language, muParser) are listed in the appendix. To rename a column right click on the name and use the Set Column As command. The special function col(x) can be used to access to the values of the column x, where x can be the column's number (as in col(2)) or its name in doublequotes (as in col("time")). This just changes a column from X to Y, etc. To put an actual name on the column, right click and choose Edit Column Description. To add a new column right click on a column name and click on either. SciDAVis will create a new plot window with the FFT amplitude curve, and a new table which contains the real part, the imaginary part, the amplitude, and the angle of the FFT. 
You can also get values from other tables using the function tablecol(t,c), where t is the table's name in doublequotes and c is the column's number or name in doublequotes (example: tablecol("Table1","time")). In this example, the amplitude curve has been normalized, and the frequencies have been shifted to obtain a centered x-scale. The variables i and j can be used to access the current row and column numbers. 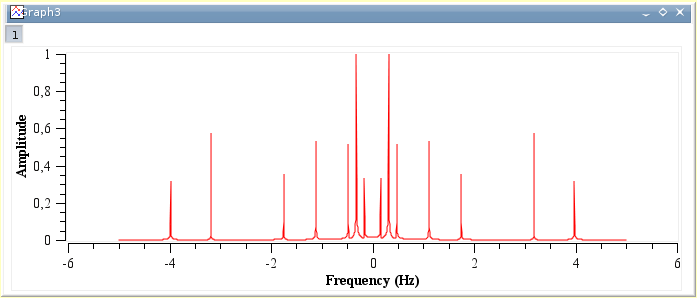
Similarly, sr and er represent the selected start and end row, respectively. There are two common ways to normalize the counts. Using Python as scripting language gives you even more possibilities, since you can not only use arbitrary Python code in the function body, but also access other objects within your project. The value of info is multiple levels (known as a nested dict). The normalized count is the count in a class divided by the total number of observations. By calling pd.jsonnormalize (jsonobj), we get: The result looks great. All nested values are flattened and converted into separate columns. If you don’t want to dig all the way down to each value use the maxlevel argument.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
